Mothers’ Global Psychological Health and Sex-specific Expression in Newborns
Stefanie R. Pilkay1*, Terri Combs-Orme2, Frances Tylavsky3, Nicole Bush4, Alicia K. Smith5
1Falk College, School of Social Work, Syracuse University, Syracuse, New York
2College of Social Work, University of Tennessee, Knoxville, Tennessee
3Department of Preventive Medicine, University of Tennesse Health and Science Center, e, Memphis, Tennessee
4Department of Psychiatry, University of California San Francisco School of Medicine, San Francisco, California
5Department of Gynecology and Obstetrics, Department of Psychiatry, Emory University School of Medicine, Atlanta, Georgia
Abstract
Summary: The prenatal environment can influence gene expression involved in the development, possibly contributing to generational patterns of psychological health. Moreover, sex-specific developmental differences in-utero may result in gene expression differences associated with the prenatal environment. However, it is not clear if maternal overall psychological symptoms will associate with newborn’s gene expression, or if such patterns are consistent between sexes. This study explored the relationships between maternal psychological health (PsyH) and newborn’s gene expression patterns. We assessed PsyH with the Brief Symptom Inventory and newborn gene expression in umbilical cord blood. We conducted combined and sex-stratified analyses of genes expressed in umbilical cord blood.
Findings: PsyH associated with differential expression of 157 genes in males. The 157 differentially expressed genes are more likely to function in metabolic processes. There were no significant differences in gene expression in females.
Application: The sex-specific nature of these findings suggests males may be more vulnerable than females to mothers’ psychological functioning during pregnancy. It is possible that the male-specific results are due in part to female newborns developing under different neuroendocrine conditions. Future research examining prenatal exposures should consider sex differences.
Introduction
Prenatal development is sensitive to maternal stress which can have lasting effects on neonate health1,2. Furthermore, males and females have shown differences in prenatal stress programming that could represent different health and developmental risks according to neonate sex3. Maternal psychological health is one maternal stress factor linked to prenatal development4 in a sex-specific manner5 that may represent sex-specific biological embedding. Maternal psychological health has been measured in different ways showing varying associations with neonate health. For example, mothers with greater affect intensity showed greater fetal motor activity during pregnancy compared to mothers with more stable affect6. Moreover, depression or anxiety during pregnancy associated with more difficult neonate temperaments7. The links between maternal psychological health and neonate health can best be explained by the process of experience influencing developmental processes, termed “biological embedding”8.
Biological Embedding
Pregnancy can be stressful for the mother9 and the additional burden of psychological symptoms can affect the fetus’ developing brain. Previous findings showed mother and newborn hypothalamus-pituitary-adrenal (HPA) axis functioning was linked to cortisol activity and suggested the HPA axis had been sensitized to stress and stimulation in neonates10 through the process of fetal programming11. Chronic stress activation can influence a sensitized HPA axis which can affect the immune system and metabolism12. These physiological adaptations increase a child’s vulnerability to environmental insults during neurodevelopment. Collectively the research suggests that women who suffer psychological health symptoms are at increased risk to have children with a greater vulnerability to stress.
Emerging evidence suggests that the mothers’ psychological health during pregnancy, can impose fetal stress that influences gene expression and fetal development13-15. Moreover, there are developmental differences between male and female embryos resulting in varying sensitivities to the prenatal environment. For example, male fetuses undergo an epigenetic process during pregnancy to masculinize the brain16. The potential for sex-specific variation in sensitivity to the prenatal environment underscores the need to examine sex-specific effects within the context of maternal psychological health. Sex-specific differences in gene expression may help explain sex-specific risk and resilience within childhood psychopathology17 as well as affective disorders found in adulthood, i.e. anxiety, bipolar disorder, depression, and post-traumatic stress disorder18.
This study aimed to determine if: 1) maternal psychological health during pregnancy will associate with gene expression in newborns, and 2) the association between maternal psychological health and newborn gene expression is sex-specific.
Results
Subjects are primarily African American mothers (African American 62% vs. Caucasian 38%) of similar age (25.04 + 5.15 vs. 28.56 + 4.61) and the newborns are comparably distributed between sexes (male [n = 61, 55%], female [n = 50, 45%]). The majority of the mothers reported some degree of PsyH symptoms (M = 50.33, SD = 7.79) that did not differ according to newborn sex (p = .18) (see Figures 1 and 2).
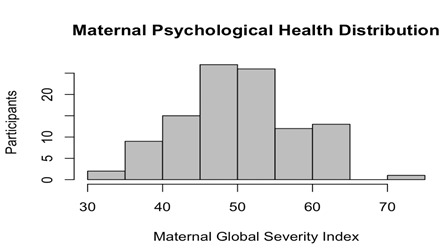
Figure 1: Histogram of Maternal Psychological Health Distribution in Full Neonate Cohort. PsyH is relatively normally distributed with a score range of 33-72 (n = 111, M = 50.33, SD = 7.8, skew = .06, kurtosis = -.32, SE = .76).
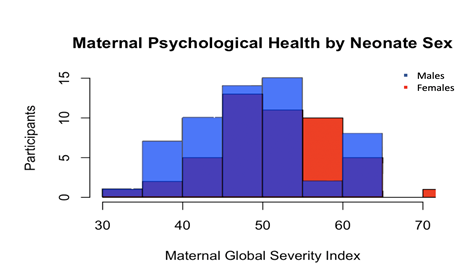
Figure 2: Maternal Psychological Health Distribution by Neonate Sex. Male and female neonates show similar distributions of maternal psychological health symptoms.
Full Cohort Associations
PsyH did not associate with newborn gene expression in the full newborn cohort while controlling for sex, race and cell composition. However, gene expression did associate with newborn sex for 17 genes, although all were located on the sex chromosomes. Plots of the effect sizes and t-statistics for the gene expression probes according to newborn sex suggests male and female newborns responded differently to PsyH (see Figure 3). Therefore, we conducted a sex-stratified analysis to investigate possible gene expression associations with PsyH unique to male or female newborns.
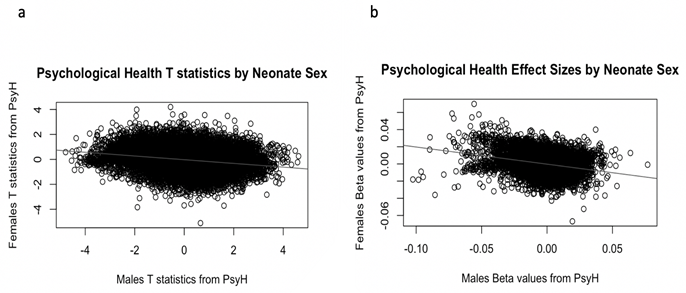
Figure 3: Plots by Newborn Sex of Association Indicators for Psychological Health. a: t-statistics show an inverse association between males and females. b: The beta values show an inverse association for gene expression and psychological health according to newborn sex.
Sex-stratified Cohort Associations
Males exhibited significant gene expression of 157 genes related to increases in PsyH scores during pregnancy (Supplemental Table 1). The 157 genes showed positive (38%) and negative (62%) associations, as modeled in a volcano plot (Figure 4), suggesting a complex interaction with the prenatal environment. These genes were enriched for a variety of metabolic functions (Table 1). For example, expression of the eukaryotic translation initiation factor 4E binding protein 2 (EIF4EBP2), a gene represented among multiple enriched biological processes is lower in males whose mothers have higher levels of PsyH symptoms (Figure 5). Female newborns showed no gene expression associations with PsyH after controlling for covariates and adjusting for multiple comparisons (FDR > .05).
Table 1. Table 1: Enrichment of Biological Processes Among Genes whose Expression associates with maternal PsyH in males.
|
Path Identifier |
Description |
# Genes |
Bayes Factor |
p-value |
|
GO:0044237 |
cellular metabolism |
46 |
7 |
.003 |
|
GO:0008152 |
metabolism |
48 |
7 |
.003 |
|
GO:0043170 |
macromolecule metabolism |
22 |
6 |
.005 |
|
GO:0044238 |
primary metabolism |
44 |
6 |
.007 |
*Adjusted p-value for multiple comparison control.
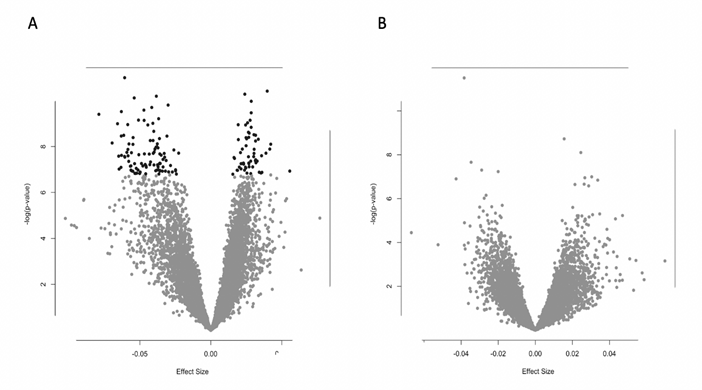
Figure 4: Volcano plot comparison of PsyH and gene expression association values by newborn sex. A: Males show more negative gene expression associations with maternal psychological health. B: Females show no gene expression associations with maternal psychological health.
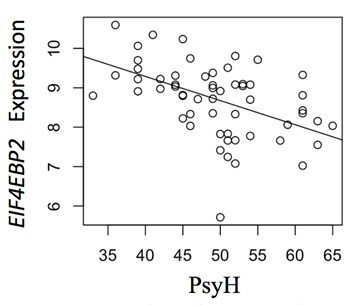
Figure 5: Maternal PsyH and EIF4EBP2 Expression in Male Newborns. EIF4EBP2 gene (ILMN_1728083) is shared among the metabolic pathways, is involved in inhibiting translation, and shows greater expression in males neonates with greater PsyH symptoms in moms. Analyses controlled for child race, cell composition, and multiple comparisons.
Discussion
Our findings suggest that maternal psychological health can influence the prenatal environment with gene expression associations unique to male newborns. These results suggest males may be more sensitive to maternal psychological health during pregnancy compared to females. Male newborns showed differential expression of 157 genes associated with greater detrimental maternal psychological health, and these genes were enriched for fundamental processes of metabolism. Moreover, a PubMed search of the 157 genes identified EIF4EBP2 as a gene of particular interest. EIF4EBP2 is involved in inhibiting translation initiation, which can affect the amount of protein produced.
Interestingly, EIF4EBP2 has been implicated in the mechanisms of Autism Spectrum Disorders in rodent models with increased ASD behaviors in mice with the gene knocked out19. Moreover, newborn expression of EIF4EBP2 has been shown to associate with maternal inflammatory responses during pregnancy20 that have been linked to maternal psychological health21. Although investigations are ongoing, EIF4EBP2 clearly plays an important role in health and the transmission of health from mother to child. The unique association for male newborns in relation to maternal psychological health, therefore, could be an indication of a sex-specific neonatal developmental response to the prenatal environment. At a minimum, our findings support previous research proposing mothers’ psychological health is associated with gene expression in male newborns that could predispose risk for or resilience to metabolic disorders22,23, enduring cognitive deficits14,15, and increased stress sensitivity5.
Maternal psychological health functioning during pregnancy should be investigated further. It is possible that some psychological health symptoms exert greater influence than others, or associate with male and female newborns differently. Future research would benefit from the inclusion of different psychological symptoms such as depression, anxiety, somatization, or combinations thereof to provide an examination of differences among symptom types. Furthermore, psychological health can influence other things such as nutrition, income, and social support which could each affect the prenatal environment.
Continuing exploration into gene expression patterns associated with prenatal variables has the potential to inform professionals seeking to foster resilience in vulnerable populations through innovative approaches to reduce risk. For example, prenatal screenings could include assessment for variables that increase the risk for adverse behavioural outcomes in offspring. Therefore, identified environmental interventions for the pregnant mother such as diet and stress reduction could be applied to foster resilience in the fetus, and later the newborn. There are many avenues to promote good psychological health through stress reduction techniques like mindfulness24, breathing exercises25, physical exercise26, and social connectedness27 to name a few. Moreover, behaviors like healthy sleep patterns28 and playing a musical instrument29 have been shown to promote positive emotions and emotion regulation that promote good maternal psychological health.
These study findings, in conjunction with previous findings5,14,15, suggest prenatal exposure to maternal psychological illness is one link for intergenerational patterns of risk, and risks may be sex-specific. Future research may deepen our understanding of mechanisms involved in these intergenerational patterns, and thereby provide new opportunities to intervene for improved quality of life. For example, this avenue of research could bolster arguments for the ongoing need for universal prenatal care, proper affordable nutrition, and other services to support mothers and families.
There are some limitations to this study. The sample is from one urban population in west Tennessee and is not generalizable to other populations. However, our replication of an association between maternal psychological health and male newborn gene expression provides additional context with previous findings5,15,15. The sample size is modest for transcriptome-wide investigations, but the transcriptome-wide analysis is an excellent tool to discover unknown or unsuspected relationships. However, analysis of the whole transcriptome significantly increases the number of conducted tests. To account for those tests, and the decreased probability of committing type I errors, we controlled for multiple comparisons with the False Discovery Rate and calculated bootstrap 95% confidence intervals to show the accuracy of estimated relationships. Lastly, we investigated these relationships in umbilical cord blood, and there may be relationships in other tissues (i.e., placenta).
The findings from our study, in conjunction with previous research30,31, suggest that social problems such as stress and psychological illness have potential to exert influence on human development across generations. Therefore, the design of interventions will benefit from biopsychosocial research, like gene expression studies, that will help illuminate pathways to risk and resilience. Knowledge like this could one day be used in screening measures to aid intervention and prevention strategies and in the design and targeting of services to those who are most in need.
Methods
Sample and Procedures
The data for this study is from the Conditions Affecting Neurocognitive Development and Learning in Early Childhood investigation (CANDLE), and the University of Tennessee Health Science Center Institutional Review Board approved all measures and procedures. This study was carried out in accordance with the Belmont Report ethical principles and guidelines for human subjects research. The sample (111 mother/infant pairs) consists of healthy mothers aged 16-40 years solicited in prenatal settings in Shelby County, Tennessee. Announcements and brochures containing information about the study were provided to all local gynecology clinics. Interested women contacted study personnel by telephone and were then screened for eligibility. Women meeting eligibility criteria were asked to visit one of two research clinics utilized for the study. Forty percent of interested women met eligibility criteria and were invited to participate. All participants signed informed consent documents, and participants under the age of eighteen years provided a parent signed informed consent. The participants provided umbilical cord blood samples immediately after birth for biological measures including gene expression. Umbilical cord blood has been used to measure newborn gene expression in multiple studies32-35.
Measures
Mothers’ psychological health (PsyH) status variable was created using the global severity index (GSI) summary score from the Brief Symptom Inventory (BSI) (Derogatis & Melisaratos, 1983). The Brief Symptom Inventory is a shortened version of the SCL-90 psychological health assessment and has been found to be sensitive to psychopathology and psychological distress36. The global severity index score has been investigated and found to be a more accurate assessment of overall psychological health functioning than the positive symptom total score in the BSI measure37.
Gene Expression
Gene expression is assessed by measuring the RNA transcript levels38. The Illumina Human WG-6 expression array was used to measure RNA transcription. Samples with less than 10% of the gene probes detected were eliminated, as well as probes with less than 10% of the samples detected within each dataset. No sample was lost based on these QC criteria. We performed quantile normalization, scaled the data, and performed a log2 transformation. Prior to analysis, updated annotation files for the Illumina Human WG-6 array were consulted for accurate gene expression measurement information39. A total of 10,821 expressed genes passed QC in these umbilical cord blood samples.
Statistical Analyses
Descriptive statistics were calculated to determine sample characteristics. We examined the association between PsyH and newborn transcriptome-wide gene expression as measured with the Illumina HumanWG-6 BeadChip. We performed multiple regression to conduct the newborn gene expression analyses and controlled for child sex, child race, and cell composition. Cell composition was estimated for each sample as previously described40. We statistically analyzed the cohort first and then conducted sex-stratified analyses controlling for child race and cell composition. Gene ontology analysis was performed on genes identified as significant using GATHER41. Bootstrap confidence intervals were generated to address potential non-normality of the data. As is standard in genetics research, we implemented the False Discovery Rate control for multiple comparisons in all gene expression analyses42.
Acknowledgements
Funding and support was provided by The Urban Child Institute and the University of Tennessee Health Science Center, with other support from the US National Institute of Child Health and Human Development, grants HD055462 and HD060713.
References
- Painter R, Osmond C, Gluckman P, et al. Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG: An International Journal of Obstetrics & Gynaecology. 2008; 115: 1243-1249.
- Almond D, Mazumder B. Health capital and the prenatal environment: the effect of Ramadan observance during pregnancy. American Economic Journal: Applied Economics. 2011; 3: 56-85.
- Glover V, Hill J. Sex differences in the programming effects of prenatal stress on psychopathology and stress responses: an evolutionary perspective. Physiology & Behavior. 2012; 106: 736-740.
- Weinstock M. The potential influence of maternal stress hormones on development and mental health of the offspring. Brain behavior and immunity. 2005; 19: 296-308.
- Mueller BR, Bale TL. Sex-Specific Programming of Offspring Emotionality Following Stress Early in Pregnancy. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2008; 28: 9055-9065. doi:10.1523/JNEUROSCI.1424-08.2008.
- DiPietro JA, Hilton SC, Hawkins M, et al. Maternal stress and affect influence fetal neurobehavioral development. Developmental psychology. 2002; 38: 659.
- Davis EP, Glynn LM, Schetter CD, et al. Prenatal Exposure to Maternal Depression and Cortisol Influences Infant Temperament. Journal of the American Academy of Child & Adolescent Psychiatry. 2007; 46: 737-746, doi:https://doi.org/10.1097/chi.0b013e318047b775.
- Hertzman C. Putting the concept of biological embedding in historical perspective. Proceedings of the National Academy of Sciences. 2012; 109: 17160-17167.
- Geller PA. Pregnancy as a Stressful Life Event. CNS Spectrums. 2004; 9: 188-197, doi:10.1017/S1092852900008981.
- Brand SR, Brennan PA, Newport DJ, et al. The impact of maternal childhood abuse on maternal and infant HPA axis function in the postpartum period. Psychoneuroendocrinology. 2010; 35: 686-693.
- Kapoor A, Dunn E, Kostaki A, et al. Fetal programming of hypothalamo-pituitary-adrenal function: prenatal stress and glucocorticoids. The Journal of physiology. 2006; 572: 31-44.
- Perry. Examining child maltreatment through a neurodevelopmental lens: Clinical applications of the neurosequential model of therapeutics. Journal of Loss and Trauma. 2009; 14: 240-255.
- Babenko O, Kovalchuk I, Metz GAS. Stress-induced perinatal and transgenerational epigenetic programming of brain development and mental health. Neuroscience and Biobehavioral Reviews. 2015; 48: 70-91. doi:http://dx.doi.org/10.1016/j.neubiorev.2014.11.013.
- Kapoor A, Kostaki A, Janus C, et al. The effects of prenatal stress on learning in adult offspring is dependent on the timing of the stressor. Behavioural Brain Research. 2009; 197: 144-149, doi:http://dx.doi.org/10.1016/j.bbr.2008.08.018.
- Lemaire V, Koehl M, Le Moal M, et al. Prenatal Stress Produces Learning Deficits Associated with an Inhibition of Neurogenesis in the Hippocampus. Proceedings of the National Academy of Sciences of the United States of America. 2000; 97: 11032-11037.
- Bale. Epigenetic and transgenerational reprogramming of brain development. Nat Rev Neurosci. 2015; 16: 332-344, doi:10.1038/nrn3818.
- Gray SA, Theall K, Lipschutz R, et al. Sex differences in the contribution of respiratory sinus arrhythmia and trauma to children’s psychopathology. Journal of psychopathology and behavioral assessment. 2017; 39: 67-78.
- Altemus M, Sarvaiya N, Epperson CN. Sex differences in anxiety and depression clinical perspectives. Frontiers in neuroendocrinology. 2014; 35: 320-330, doi:10.1016/j.yfrne.2014.05.004.
- Gkogkas CG, Khoutorsky A, Ran I, et al. Autism-related deficits via dysregulated eIF4E-dependent translational control. Nature. 2013; 493: 371.
- Zhou J, Zhang X, Zhang H, et al. Use of data mining to determine changes in the gene expression profiles of rat embryos following prenatal exposure to inflammatory stimulants. Molecular medicine reports. 2013; 8: 95-102.
- Christian LM. Stress and Immune Function during Pregnancy: An Emerging Focus in Mind-Body Medicine. Curr Dir Psychol Sci. 2015; 24: 3-9, doi:10.1177/0963721414550704.
- Harris A, Seckl J. Glucocorticoids, prenatal stress and the programming of disease. Hormones and Behavior. 2011; 59: 279-289, doi:https://doi.org/10.1016/j.yhbeh.2010.06.007.
- Tamashiro KLK, Moran TH. Perinatal environment and its influences on metabolic programming of offspring. Physiology & Behavior. 2010; 100: 560-566, doi:https://doi.org/10.1016/j.physbeh.2010.04.008.
- Khoury B, Sharma M, Rush SE, et al. Mindfulness-based stress reduction for healthy individuals: A meta-analysis. Journal of psychosomatic research. 2015; 78: 519-528.
- Varvogli L, Darviri C. Stress Management Techniques: evidence-based procedures that reduce stress and promote health. Health science journal. 2011; 5-74.
- Yau SY, Lau BWM, So KF. Adult hippocampal neurogenesis: a possible way how physical exercise counteracts stress. Cell Transplantation. 2011; 20: 99-111.
- Haslam C, Cruwys T, Haslam SA, et al. Social connectedness and health. Encyclopaedia of geropsychology. 2015; 46-41.
- Shen L, van Schie J, Ditchburn G, et al. Positive and negative emotions: Differential associations with sleep duration and quality in adolescents. Journal of youth and adolescence. 2018; 47: 2584-2595.
- Moore KS. A systematic review on the neural effects of music on emotion regulation: implications for music therapy practice. Journal of music therapy. 2013; 50: 198-242.
- Meaney MJ. Maternal care, gene expression, and the transmission of individual differences in stress reactivity across generations. Annual review of neuroscience. 2001; 24: 1161-1192.
- Yehuda R, Bierer LM. Transgenerational transmission of cortisol and PTSD risk. Progress in brain research. 2007; 167: 121-135.
- Adkins RM, Krushkal J, Tylavsky FA, et al. Racial differences in gene-specific DNA methylation levels are present at birth. Birth Defects Research Part A: Clinical and Molecular Teratology. 2011; 91: 728-736.
- Adkins RM, Thomas F, Tylavsky FA, et al. Parental ages and levels of DNA methylation in the newborn are correlated. BMC medical genetics. 2011; 12: 47.
- Adkins RM, Tylavsky FA, Krushkal J. Newborn umbilical cord blood DNA methylation and gene expression levels exhibit limited association with birth weight. Chemistry & biodiversity. 2012; 9: 888-899.
- Krushkal J, et al. Epigenetic analysis of neurocognitive development at 1 year of age in a community-based pregnancy cohort. Behavior genetics. 2014; 44: 113-125.
- Sitarenios G, Kovacs M, Maruish M. The use of psychological testing for treatment planning and outcomes assessment. Use of the Children's Depression Inventory. 1999; 267-298.
- Derogatis LR, Melisaratos N. The brief symptom inventory: an introductory report. Psychological medicine. 1983; 13: 595-605.
- Marioni JC, Mason CE, Mane SM, et al. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome research. 2008; 18: 1509-1517.
- Arloth J, Bader DM, Röh S, et al. Re-Annotator: Annotation Pipeline for Microarray Probe Sequences. PloS one. 2015; 10: e0139516, doi:10.1371/journal.pone.0139516.
- Mozhui K, Smith AK, Tylavsky FA. Ancestry dependent DNA methylation and influence of maternal nutrition. PloS one. 2015; 10: e0118466.
- Chang JT, Nevins JR. GATHER: a systems approach to interpreting genomic signatures. Bioinformatics (Oxford, England). 2006; 22: 2926-2933, doi:10.1093/bioinformatics/btl483.
- Storey JD, Taylor JE, Siegmund D. Strong control, conservative point estimation and simultaneous conservative consistency of false discovery rates: a unified approach. Journal of the Royal Statistical Society: Series B (Statistical Methodology). 2004; 66: 187-205.
Supplemental Table.
Genes differentially expressed in male newborns according to maternal PsyH
|
Probe ID |
Gene |
Beta Coefficient |
Bootstrap 95% CI |
FDR adjusted p-value |
|
Probe ID |
Gene |
Beta Coefficient |
Bootstrap 95% CI |
FDR adjusted p-value |
|||||||||
|
ILMN_2185264 |
ZNF461 |
-0.078886042 |
-.119, -.038 |
0.039400571 |
|
ILMN_2126802 |
RPS27L |
-0.054882677 |
-.084, -.029 |
0.044812759 |
|||||||||
|
ILMN_1762573 |
LOC401630 |
-0.069570218 |
-.108, -.035 |
0.044812759 |
|
ILMN_2058841 |
LILRA6 |
-0.054787834 |
-.083, -.025 |
0.043364783 |
|||||||||
|
ILMN_2096442 |
LOC260339 |
-0.065750621 |
-.097, -.032 |
0.039400571 |
|
ILMN_2380101 |
PHACTR4 |
-0.053983378 |
-.070, -.030 |
0.039400571 |
|||||||||
|
ILMN_1746917 |
LOC729843 |
-0.064957403 |
-.084, -.038 |
0.045569436 |
|
ILMN_2190850 |
PPID |
-0.053745272 |
-.086, -.025 |
0.047760063 |
|||||||||
|
ILMN_2175737 |
ZNF826 |
-0.064650594 |
-.099, -.023 |
0.048165878 |
|
ILMN_2179579 |
SNHG3 |
-0.053519627 |
-.085, -.024 |
0.049176039 |
|||||||||
|
ILMN_2198823 |
H6PD |
-0.063303215 |
-.098, -.031 |
0.047760063 |
|
ILMN_2106002 |
ACBD7 |
-0.052888042 |
-.085, -.027 |
0.049176039 |
|||||||||
|
ILMN_2404320 |
SNTN |
-0.06312612 |
-.095, -.033 |
0.043364783 |
|
ILMN_2395496 |
KLK7 |
-0.052570497 |
-.087, -.020 |
0.048165878 |
|||||||||
|
ILMN_1698766 |
PYCARD |
-0.063042137 |
-.082, -.041 |
0.039400571 |
|
ILMN_2115011 |
FGD2 |
-0.052162454 |
-.082, -.024 |
0.049176039 |
|||||||||
|
ILMN_2141523 |
MRPL44 |
-0.06294615 |
-.097, -.029 |
0.045569436 |
|
ILMN_2084489 |
ZNF595 |
-0.051316734 |
-.082, -.019 |
0.048165878 |
|||||||||
|
ILMN_2402499 |
SC4MOL |
-0.06135017 |
-.089, -.035 |
0.043364783 |
|
ILMN_2215965 |
CYP2B6 |
-0.050994618 |
-.074, -.024 |
0.039400571 |
|||||||||
|
ILMN_2203876 |
CCDC68 |
-0.060809072 |
-.094, -.027 |
0.045569436 |
|
ILMN_1712357 |
HNRPK |
-0.050745076 |
-.074, -.025 |
0.045569436 |
|||||||||
|
ILMN_1876838 |
|
-0.060756232 |
-.088, -.039 |
0.039400571 |
|
ILMN_1757914 |
C19orf56 |
-0.050123353 |
-.072, -.027 |
0.049176039 |
|||||||||
|
ILMN_1715635 |
ATP6V0E1 |
-0.060589336 |
-.095, -.033 |
0.045569436 |
|
ILMN_1741491 |
ZNHIT1 |
-0.049742067 |
-.083, -.026 |
0.048871645 |
|||||||||
|
ILMN_2357377 |
TERF1 |
-0.059190782 |
-.091, -.027 |
0.045569436 |
|
ILMN_2178186 |
PIGW |
-0.04887585 |
-.076, -.022 |
0.047760063 |
|||||||||
|
ILMN_2208491 |
RPLP0P2 |
-0.058546721 |
-.089, -.036 |
0.039400571 |
|
ILMN_2127416 |
GSR |
-0.047914114 |
-.073, -.028 |
0.047760063 |
|||||||||
|
ILMN_2066249 |
RPP30 |
-0.058426512 |
-.095, -.031 |
0.045569436 |
|
ILMN_1671494 |
USP5 |
-0.047160548 |
-.068, -.032 |
0.039400571 |
|||||||||
|
ILMN_2217955 |
TTC21B |
-0.058323114 |
-.091, -.029 |
0.045569436 |
|
ILMN_1689710 |
C16orf50 |
-0.047007495 |
-.069, -.026 |
0.039400571 |
|||||||||
|
ILMN_2281089 |
STEAP3 |
-0.057859572 |
-.092, -.025 |
0.047760063 |
|
ILMN_2102580 |
UTP20 |
-0.046813344 |
-.071, -.021 |
0.045569436 |
|||||||||
|
ILMN_1679809 |
GSTP1 |
-0.057681223 |
-.088, -.031 |
0.044812759 |
|
ILMN_2409720 |
SLA2 |
-0.04625212 |
-.070, -.018 |
0.047760063 |
|||||||||
|
ILMN_1651358 |
HBE1 |
-0.056699663 |
-.081, -.020 |
0.047760063 |
|
ILMN_2245686 |
GYG2 |
-0.045899964 |
-.073, -.018 |
0.049176039 |
|||||||||
|
ILMN_2346562 |
ZNF273 |
-0.056319333 |
-.090, -.029 |
0.048165878 |
|
ILMN_1765621 |
HDGF |
-0.045702693 |
-.068, -.020 |
0.047760063 |
|||||||||
|
ILMN_2382657 |
ARHGAP9 |
-0.056298906 |
-.086, -.027 |
0.047760063 |
|
ILMN_1917044 |
|
-0.045493069 |
-.072, -.030 |
0.047760063 |
|||||||||
|
ILMN_2049228 |
NUDT4P1 |
-0.055937291 |
-.086, -.028 |
0.045569436 |
|
ILMN_1756942 |
SP3 |
-0.044384942 |
-.056, -.025 |
0.039400571 |
|||||||||
|
ILMN_2180997 |
GTF2IRD2B |
-0.055805828 |
-.088, -.031 |
0.048165878 |
|
ILMN_2151168 |
SLC30A6 |
-0.043380056 |
-.065, -.019 |
0.045569436 |
|||||||||
|
ILMN_1728083 |
EIF4EBP2 |
-0.054931752 |
-.080, -.032
|
0.048165878 |
|
ILMN_2378670 |
SNX15 |
-0.043006845 |
-.067, -.016 |
0.047760063 |
|||||||||
|
ILMN_2070477 |
TAF8 |
-0.042691067 |
-.064, -.020 |
0.043364783 |
|
ILMN_2137464 |
DVL3 |
-0.035973219 |
-.055, -.017 |
0.047760063 |
|||||||||
|
ILMN_1715698 |
MGC71993 |
-0.042485825 |
-.064, -.020 |
0.047760063 |
|
ILMN_2145143 |
FKBP9 |
-0.035961146 |
-.052, -.012 |
0.045569436 |
|||||||||
|
ILMN_2292696 |
COX15 |
-0.041623601 |
-.062, -.028 |
0.039400571 |
|
ILMN_2331658 |
C3orf17 |
-0.03585831 |
-.054, -.017 |
0.048165878 |
|||||||||
|
ILMN_1742400 |
CEP350 |
-0.040846365 |
-.054, -.022 |
0.039400571 |
|
ILMN_1794522 |
EIF5A |
-0.035854562 |
-.061, -.019 |
0.048165878 |
|||||||||
|
ILMN_2280441 |
PACRG |
-0.040702491 |
-.059, -.017 |
0.043364783 |
|
ILMN_2141030 |
LOC641522 |
-0.035638955 |
-.054, -.014 |
0.047760063 |
|||||||||
|
ILMN_2042941 |
TMEM159 |
-0.040579203 |
-.059, -.019 |
0.045569436 |
|
ILMN_1797964 |
ARL6IP6 |
-0.024273555 |
-.030, -.004 |
0.049176039 |
|||||||||
|
ILMN_1758100 |
GALR3 |
-0.040217106 |
-.060, -.019 |
0.043364783 |
|
ILMN_1773850 |
FXC1 |
-0.022685488 |
-.035, -.011 |
0.045569436 |
|||||||||
|
ILMN_1739792 |
RHOG |
-0.040075792 |
-.061, -.018 |
0.047760063 |
|
ILMN_1682736 |
LOC643452 |
0.01550105 |
.004, .024 |
0.049177556 |
|||||||||
|
ILMN_2410362 |
ACBD5 |
-0.039916622 |
-.059, -.018 |
0.045569436 |
|
ILMN_1675852 |
LOC650518 |
0.016336257 |
.008, .026 |
0.04559356 |
|||||||||
|
ILMN_2055271 |
A1BG |
-0.039415396 |
-.056, -.017 |
0.045569436 |
|
ILMN_1721713 |
EXOSC9 |
0.016725851 |
.006, .025 |
0.04681483 |
|||||||||
|
ILMN_2358652 |
NXF1 |
-0.039356826 |
-.062, -.019 |
0.045569436 |
|
ILMN_1659523 |
USP39 |
0.018544909 |
.003, .024 |
0.048165878 |
|||||||||
|
ILMN_2374383 |
TSPAN17 |
-0.038850669 |
-.059, -.021 |
0.048165878 |
|
ILMN_1776347 |
TCP1 |
0.018823076 |
.009, .026 |
0.048165878 |
|||||||||
|
ILMN_2162972 |
LYZ |
-0.03863507 |
-.055, -.016 |
0.045569436 |
|
ILMN_1767992 |
SLC12A6 |
0.019306512 |
.009, .029 |
0.039400571 |
|||||||||
|
ILMN_1651506 |
NCOA6IP |
-0.038395972 |
-.049, -.019 |
0.039400571 |
|
ILMN_1704206 |
NPSR1 |
0.019418619 |
.0003, .024 |
0.043364783 |
|||||||||
|
ILMN_1682938 |
ARF3 |
-0.038045144 |
-.060, -.014 |
0.048165878 |
|
ILMN_2192683 |
DHX37 |
0.020070541 |
.007, .026 |
0.047760063 |
|||||||||
|
ILMN_2277252 |
PPFIBP1 |
-0.038031935 |
-.055, -.017 |
0.045569436 |
|
ILMN_1662896 |
BRWD2 |
0.0209141 |
.009, .030 |
0.045569436 |
|||||||||
|
ILMN_2178201 |
ZNF43 |
-0.037734631 |
-.055, -.018 |
0.044812759 |
|
ILMN_1776147 |
C21orf59 |
0.021299857 |
.008, .031 |
0.048165878 |
|||||||||
|
ILMN_2255142 |
TRIM34 |
-0.037587268 |
-.057, -.013 |
0.047760063 |
|
ILMN_1727761 |
GMEB1 |
0.021302431 |
.006, .026 |
0.047760063 |
|||||||||
|
ILMN_2115974 |
GSDM1 |
-0.037211655 |
-.055, -.014 |
0.045569436 |
|
ILMN_1693421 |
RPN2 |
0.022035022 |
.010, .032 |
0.047760063 |
|||||||||
|
ILMN_1660869 |
LOC643438 |
-0.037161222 |
-.056, -.020 |
0.047760063 |
|
ILMN_1725169 |
INTS12 |
0.022214042 |
.005, .034 |
0.045569436 |
|||||||||
|
ILMN_1750805 |
ARHGAP30 |
-0.036936857 |
-.053, -.023 |
0.048165878 |
|
ILMN_1737413 |
MSH2 |
0.023029546 |
.005, .030 |
0.047760063 |
|||||||||
|
ILMN_2261600 |
FCGR1B |
-0.036794728 |
-.046, -.017 |
0.039400571 |
|
ILMN_1916094 |
|
0.023133907 |
.009, .035 |
0.047760063 |
|||||||||
|
ILMN_1684434 |
SLC17A5 |
-0.036576672 |
-.057, -.014 |
0.049176039 |
|
ILMN_1677376 |
CHD7 |
0.023791824 |
.012, .037 |
0.048165878 |
|||||||||
|
ILMN_2190851 |
PPID |
-0.036289407 |
-.052, -.014 |
0.047760063 |
|
ILMN_1774974 |
CLUAP1 |
0.023952943 |
.011, .030 |
0.039400571 |
|||||||||
|
ILMN_2196232 |
C1orf210 |
-0.036197628 |
-.052, -.016 |
0.043364783 |
|
ILMN_1748018 |
GORASP2 |
0.02441163 |
.010, .032 |
0.047760063 |
|||||||||
|
ILMN_2359096 |
SS18 |
-0.034992783 |
-.053, -.015 |
0.045569436 |
|
ILMN_1801833 |
ARHGAP24 |
0.024525524 |
.015, .038 |
0.039400571 |
|||||||||
|
ILMN_2252136 |
YWHAE |
-0.034150242 |
-.060, -.021 |
0.048165878 |
|
ILMN_1771801 |
SIRPG |
0.024716906 |
.012, .036 |
0.048165878 |
|||||||||
|
ILMN_2406532 |
F11R |
-0.033700911 |
-.051, -.011 |
0.047760063 |
|
ILMN_1684724 |
CR2 |
0.024839519 |
.012, .037 |
0.043364783 |
|||||||||
|
ILMN_2263236 |
HFE |
-0.033652648 |
-.053, -.014 |
0.047760063 |
|
ILMN_2136133 |
PABPC1 |
0.024864516 |
.010, .033 |
0.045569436 |
|||||||||
|
ILMN_2299795 |
CPM |
-0.032837325 |
-.045, -.010 |
0.045569436 |
|
ILMN_1720270 |
CDR2 |
0.025226775 |
.006, .032 |
0.043364783 |
|||||||||
|
ILMN_1728957 |
ANKRD5 |
-0.032553034 |
-.049, -.012 |
0.045569436 |
|
ILMN_1789653 |
PBLD |
0.025480103 |
.010, .038 |
0.049176039 |
|||||||||
|
ILMN_1775919 |
C6orf79 |
-0.031994525 |
-.049, -.013 |
0.048165878 |
|
ILMN_2048822 |
NUDCD2 |
0.025886067 |
.013, .033 |
0.039400571 |
|||||||||
|
ILMN_2323302 |
SON |
-0.031503972 |
-.051, -.018 |
0.047760063 |
|
ILMN_1655625 |
GPATCH1 |
0.026500147 |
.007, .041 |
0.043364783 |
|||||||||
|
ILMN_1810488 |
NFYC |
-0.030915679 |
-.044, -.012 |
0.043364783 |
|
ILMN_1725175 |
FOSL2 |
0.02654905 |
.010, .039 |
0.045569436 |
|||||||||
|
ILMN_1712400 |
SERPINB6 |
-0.030091528 |
-.042, -.017 |
0.039400571 |
|
ILMN_1749586 |
LOC642914 |
0.026746809 |
.016, .046 |
0.047760063 |
|||||||||
|
ILMN_1770673 |
AKNA |
-0.029749607 |
-.043, -.009 |
0.048871645 |
|
ILMN_1717852 |
USH1G |
0.026754287 |
.016, .039 |
0.043364783 |
|||||||||
|
ILMN_2144116 |
CPSF2 |
-0.02838741 |
-.040, -.012 |
0.047760063 |
|
ILMN_2156953 |
ZFAND6 |
0.026820464 |
.015, .036 |
0.047760063 |
|||||||||
|
ILMN_1730879 |
CBY1 |
-0.027131446 |
-.039, -.014 |
0.048871645 |
|
ILMN_2186482 |
TMED7 |
0.026990153 |
.015, .043 |
0.047760063 |
|||||||||
|
ILMN_1744113 |
TNFAIP8L2 |
-0.026052743 |
-.034, -.008 |
0.045569436 |
|
ILMN_1672446 |
RPL11 |
0.027072111 |
.008, .040 |
0.043364783 |
|||||||||
|
ILMN_2122022 |
ZNF639 |
-0.024844573 |
-.038, -.010 |
0.048165878 |
|
ILMN_1704956 |
SMTNL1 |
0.027710644 |
.007, .039 |
0.039400571 |
|||||||||
|
ILMN_1807649 |
SPOPL |
-0.024839798 |
-.029, -.007 |
0.048165878 |
|
ILMN_1818935 |
|
0.028159219 |
.016, .044 |
0.047760063 |
|||||||||
|
ILMN_1692535 |
DPP4 |
0.028160224 |
.012, .036 |
0.049176039 |
|
ILMN_2130525 |
TSPAN13 |
0.031815095 |
.015, .043 |
0.047760063 |
|||||||||
|
ILMN_2114876 |
RPL11 |
0.028446346 |
.009, .043 |
0.04180435 |
|
ILMN_2381064 |
TPD52 |
0.032692639 |
.017, .050 |
0.047760063 |
|||||||||
|
ILMN_1764323 |
LOC124512 |
0.028469498 |
.016, .040 |
0.039400571 |
|
ILMN_1652085 |
MPHOSPH10 |
0.033498448 |
.017, .048 |
0.043364783 |
|||||||||
|
ILMN_2410771 |
KEAP1 |
0.028499536 |
.008, .033 |
0.039400571 |
|
ILMN_1657873 |
XPO4 |
0.034105854 |
.017, .054 |
0.048871645 |
|||||||||
|
ILMN_1853160 |
|
0.029498432 |
.017, .044 |
0.047760063 |
|
ILMN_2200636 |
KIAA1267 |
0.035372831 |
.017, .048 |
0.049176039 |
|||||||||
|
ILMN_1653129 |
CSTF2 |
0.030208328 |
.013, .039 |
0.045569436 |
|
ILMN_1909223 |
|
0.035757529 |
.017, .050 |
0.045569436 |
|||||||||
|
ILMN_1730791 |
LOC646783 |
0.030379729 |
.015, .043 |
0.043364783 |
|
ILMN_2103774 |
PIP5KL1 |
0.036060422 |
.014, .047 |
0.048871645 |
|||||||||
|
ILMN_2151048 |
STAG1 |
0.030436997 |
.019, .041 |
0.045569436 |
|
ILMN_1837286 |
|
0.038877769 |
.019, .055 |
0.045569436 |
|||||||||
|
ILMN_1880113 |
|
0.030540174 |
.013, .045 |
0.045569436 |
|
ILMN_1819251 |
|
0.039768665 |
.019, .053 |
0.039400571 |
|||||||||
|
ILMN_1879078 |
|
0.031327435 |
.013, .047 |
0.047760063 |
|
ILMN_2379788 |
HIF1A |
0.041570256 |
.028, .062 |
0.045569436 |
|||||||||
|
ILMN_1888252 |
|
0.031513924 |
.015, .048 |
0.043364783 |
|
ILMN_1798874 |
TMEM85 |
0.042248923 |
.018, .057 |
0.044812759 |
|||||||||
|
ILMN_1662845 |
NBPF11 |
0.031580898 |
.018, .049 |
0.043364783 |
|
ILMN_2379762 |
NPM1 |
0.055644091 |
.027, .087 |
0.048165878 |
|||||||||
|
ILMN_1748141 |
AMOTL1 |
0.031599523 |
.015, .048 |
0.047760063 |
|
|
|
|
|
|
|||||||||
